Abstract
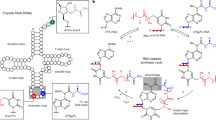
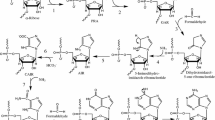
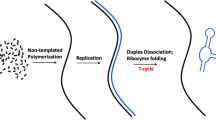
The central dogma of molecular biology dictates that, with only a few exceptions, information proceeds from DNA to protein through an RNA intermediate. Examining the enigmatic steps from prebiotic to biological chemistry, we take another road suggesting that primordial peptides acted as template for the self-assembly of the first nucleic acids polymers. Arguing in favour of a sort of archaic “reverse translation” from proteins to RNA, our basic premise is a Hadean Earth where key biomolecules such as amino acids, polypeptides, purines, pyrimidines, nucleosides and nucleotides were available under different prebiotically plausible conditions, including meteorites delivery, shallow ponds and hydrothermal vents scenarios. Supporting a protein-first scenario alternative to the RNA world hypothesis, we propose the primeval occurrence of short two-dimensional peptides termed “selective amino acid- and nucleotide-matching oligopeptides” (henceforward SANMAOs) that noncovalently bind at the same time the polymerized amino acids and the single nucleotides dispersed in the prebiotic milieu. In this theoretical paper, we describe the chemical features of this hypothetical oligopeptide, its biological plausibility and its virtues from an evolutionary perspective. We provide a theoretical example of SANMAO’s selective pairing between amino acids and nucleosides, simulating a poly-Glycine peptide that acts as a template to build a purinic chain corresponding to the glycine’s extant triplet codon GGG. Further, we discuss how SANMAO might have endorsed the formation of low-fidelity RNA’s polymerized strains, well before the appearance of the accurate genetic material’s transmission ensured by the current translation apparatus.


Similar content being viewed by others
Data Availability
All data and materials generated or analyzed during this study are included in the manuscript. The authors had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
References
Apostolopoulos V, Bojarska J, Chai T-T, Elnagdy S, Kaczmarek K et al (2021) A global review on short peptides: frontiers and perspectives. Molecules 26(2):430. https://doi.org/10.3390/molecules26020430
Baaske P, Weinert F, Duhr S, Lemke K, Russell MJ, Braun D (2007) Extreme accumulation of nucleotides in simulated hydrothermal pore systems. Proc Natl Acad Sci USA 104:9346–9351
Becker S, Feldmann J, Wiedemann S, Okamura H, Schneider C et al (2019) Unified prebiotically plausible synthesis of pyrimidine and purine RNA ribonucleotides. Science 366(6461):76–82. https://doi.org/10.1126/science.aax2747
Bernheim A, Millman A, Ofir G et al (2021) Prokaryotic viperins produce diverse antiviral molecules. Nature 589:120–124. https://doi.org/10.1038/s41586-020-2762-2
Bhowmik S, Krishnamurthy R (2019) The role of sugar-backbone heterogeneity and chimeras in the simultaneous emergence of RNA and DNA. Nat Chem 11:1009–1018
Biro JC (2007) The Proteomic Code: a molecular recognition code for proteins. Theor Biol Med Model 4:45. https://doi.org/10.1186/1742-4682-4-45
Boyd ES, Amenabar MJ, Poudel S, Templeton AS (2020) Bioenergetic constraints on the origin of autotrophic metabolism. Philos Trans A 378(2165):20190151. https://doi.org/10.1098/rsta.2019.0151. (published online 6 Jan 2020)
Broadley MW, Bekaert DV, Piani L et al (2022) Origin of life-forming volatile elements in the inner Solar System. Nature 611:245–255. https://doi.org/10.1038/s41586-022-05276-x
Carter CW Jr, Kraut J (1973) A proposed model for interaction of polypeptides with RNA. Proc Natl Acad Sci USA 71(2):283–287. https://doi.org/10.1073/pnas.71.2.283
Cartwright JH, Russell MJ (2019) The origin of life: the submarine alkaline vent theory at 30. Interface Focus. https://doi.org/10.1098/rsfs.2019.0104
Chandramouly G, Zhao J, McDevitt S, Rusanov T, Hoang T et al (2021) Polθ reverse transcribes RNA and promotes RNA-templated DNA repair. Sci Adv 7(24):eabf1771. https://doi.org/10.1126/sciadv.abf1771. (published online 11 June 2021)
Damer B, Deamer D (2020) The hot spring hypothesis for an origin of life. Astrobiology. https://doi.org/10.1089/ast.2019.2045
Di Giulio M (1997) On the RNA world: evidence in favor of an early ribonucleopeptide world. J Mol Evol 45:571–578
Di Giulio M (2015) A model for the origin of the first mRNAs. J Mol Evol 81:10–17
Dishman AF, Tyler RC, Fox JC, Kleist AB, Prehoda KE et al (2021) Evolution of fold switching in a metamorphic protein. Science 371(6524):86–90. https://doi.org/10.1126/science.abd8700
Eigenbrode JL, Summons RE, Steele A, Freissinet C, Millan M et al (2018) Organic matter preserved in 3-billion-year-old mudstones at Gale crater, Mars. Science 360(6393):1096–1101. https://doi.org/10.1126/science.aas9185
Ferro K, Peuß R, Yang W, Rosenstiel P, Schulenburg H, Kurtz J (2019) Experimental evolution of immunological specificity. Proc Natl Acad Sci USA 116(41):20598–20604. https://doi.org/10.1073/pnas.1904828116
Franz LP, Douzi B, Durand E, Dyer DH, Voulhoux R, Forest KT (2011) Structure of the minor pseudopilin XcpW from the Pseudomonas aeruginosa type II secretion system. Acta Crystallogr D 67:124–130
Frenkel-Pinter M, Haynes JW, Martin C, Petrov AS, Burcar BT et al (2019) Selective incorporation of proteinaceous over nonproteinaceous cationic amino acids in model prebiotic oligomerization reactions. Proc Natl Acad Sci USA 116(33):16338–16346. https://doi.org/10.1073/pnas.1904849116
Frenkel-Pinter M, Haynes JW, Mohyeldin AM et al (2020) Mutually stabilizing interactions between proto-peptides and RNA. Nat Commun 11:3137
Fryer P (2012) Serpentinite mud volcanism: observations, processes, and implications. Annu Rev Mar Sci 4:345–373. https://doi.org/10.1146/annurev-marine-120710-100922
Fusz S, Eisenführ A, Srivatsan SG, Heckel A, Famulok M (2005) A ribozyme for the aldol reaction. Chem Biol 12(8):941–950. https://doi.org/10.1016/j.chembiol.2005.06.008
Gilbert W (1986) Origin of life: the RNA world. Nature 319:618. https://doi.org/10.1038/319618a0
Holm NG, Dumont M, Ivarsson M, Konn C (2006) Alkaline fluid circulation in ultramafic rocks and formation of nucleotide constituents: a hypothesis. Geochem Trans 7:7
Holm NG, Oze C, Mousis O, Waite JH, Guilbert-Lepoutre A (2015) Serpentinization and the formation of H2 and CH4 on celestial bodies (planets, moons, comets). Astrobiology 15(7):587–600. https://doi.org/10.1089/ast.2014.1188
Hutchison CA III, Chuang R-Y, Noskov VN, Assad-Garcia N, Deerinc TJ et al (2016) Design and synthesis of a minimal bacterial genome. Science. https://doi.org/10.1126/science.aad6253
Jacobs WM (2021) Self-assembly of biomolecular condensates with shared components. Phys Rev Lett 126:258101 (Published 24 June 2021)
Jerome CA, Kim H-J, Mojzsis SJ, Benner SA, Biondi E (2022) Catalytic synthesis of polyribonucleic acid on prebiotic rock glasses. Astrobiology 22(6):629–636. https://doi.org/10.1089/ast.2022.0027
Kempes CP, Krakauer DC (2021) The multiple paths to multiple life. J Mol Evol 89:415–426
Kirschning A (2021) The coenzyme/protein pair and the molecular evolution of life. Nat Prod Rep 38(5):993–1010. https://doi.org/10.1039/d0np00037j
Kitadai N, Maruyama S (2018) Origins of building blocks of life: a review. Geosci Front 9(4):1117–1153
Krasnokutski SA, Chuang K-J, Jäger C, Ueberschaar N, Henning TH (2022) A pathway to peptides in space through the condensation of atomic carbon. Nat Astron 6:381–386
Kroiss D, Ashkenasy G, Braunschweig AB, Tuttle T, Ulijn RV (2019) Catalyst: can systems chemistry unravel the mysteries of the chemical origins of life? Chem 5:1917–1920
Lamadrid HM, Donald Rimstidt J, Schwarzenbach EM, Klein F, Ulrich S et al (2017) Effect of water activity on rates of serpentinization of olivine. Nat Commun 8:16107
Lella M, Mahalakshmi R (2017) Metamorphic proteins: emergence of dual protein folds from one primary sequence. Biochemistry 56(24):2971–2984. https://doi.org/10.1021/acs.biochem.7b00375. (Publication Date 1 June 2017)
Lundin D, Berggren G, Logan DT, Sjöberg B-M (2015) The origin and evolution of ribonucleotide reduction. Life 5(1):604–636. https://doi.org/10.3390/life5010604
McMullen A, Muñoz Basagoiti M, Zeravcic Z et al (2022) Self-assembly of emulsion droplets through programmable folding. Nature 610:502–506
Ménez B, Pisapia C, Andreani M, Jamme F, Vanbellingen QP et al (2018) Abiotic synthesis of amino acids in the recesses of the oceanic lithosphere. Nature 564(7734):59–63. https://doi.org/10.1038/s41586-018-0684-z. (Epub 7 Nov 2018)
Michel JA, Yunker PJ (2019) Structural hierarchy confers error tolerance in biological materials. Proc Natl Acad Sci USA 116(8):2875–2880. https://doi.org/10.1073/pnas.1813801116
Milner-White EJ (2019) Protein three-dimensional structures at the origin of life. Interface Focus 9(6):20190057. https://doi.org/10.1098/rsfs.2019.0057(Publishedonline18Oct2019)
Milner-White EJ, Russell MJ (2008) Predicting the conformations of peptides and proteins in early evolution. Biol Direct 3:3. https://doi.org/10.1186/1745-6150-3-3
Mossela E, Steel M (2005) Random biochemical networks: the probability of self-sustaining autocatalysis. J Theor Biol 233(3):327–336. https://doi.org/10.1016/j.jtbi.2004.10.011
Müller F, Escobar L, Xu F, Węgrzyn E, Nainytė M et al (2022) A prebiotically plausible scenario of an RNA–peptide world. Nature 605:279–284
Oba Y, Takano Y, Furukawa Y, Koga T, Glavin DP et al (2022) Identifying the wide diversity of extraterrestrial purine and pyrimidine nucleobases in carbonaceous meteorites. Nat Commun 13:2008
Otten R, Pádua RAP, Adrian Bunzel H, Nguyen V, Pitsawong W et al (2020) How directed evolution reshapes the energy landscape in an enzyme to boost catalysis. Science 370(6523):1442–1446. https://doi.org/10.1126/science.abd3623
Pang TY, Lercher MJ (2018) Each of 3,323 metabolic innovations in the evolution of E. coli arose through the horizontal transfer of a single DNA segment. Proc Natl Acad Sci USA 116(1):187–192. https://doi.org/10.1073/pnas.1718997115
Postec A, Quéméneur M, Bes M, Mei N, Benaïssa F et al (2015) Microbial diversity in a submarine carbonate edifice from the serpentinizing hydrothermal system of the Prony Bay (New Caledonia) over a 6-year period. Front Microbiol. https://doi.org/10.3389/fmicb.2015.00857
Preiner M, Xavier JC, Sousa FL, Zimorski V, Neubeck A et al (2018) Serpentinization: connecting geochemistry, ancient metabolism and industrial hydrogenation. Life 8(4):41. https://doi.org/10.3390/life8040041
Root-Bernstein RS (1982) Amino acid pairing. J Theor Biol 94(4):885–894. https://doi.org/10.1016/0022-5193(82)90083-2
Rouméjon S, Cannat M (2014) Serpentinization of mantle-derived peridotites at mid-ocean ridges: mesh texture development in the context of tectonic exhumation. Geochem Geophys Geosyst 15(6):2354–2379. https://doi.org/10.1002/2013GC005148Citations:55. (First published 23 May 2014)
Russell MJ, Ponce A (2020) Six ‘Must-Have’ minerals for life’s emergence: olivine, pyrrhotite, bridgmanite, serpentine, fougerite and mackinawite. Life 10(11):291. https://doi.org/10.3390/life10110291
Russell MJ, Hall AJ, Martin W (2010) Serpentinization as a source of energy at the origin of life. Geobiology 8(5):355–371. https://doi.org/10.1111/j.1472-4669.2010.00249.x
Schiller MR (2016) The minimotif synthesis hypothesis for the origin of life. J Transl Sci 2(5):289–296. https://doi.org/10.15761/JTS.1000154(Publishedonline19July2016)
Schneider C, Becker S, Okamura H, Crisp A, Amatov T et al (2018) Noncanonical RNA nucleosides as molecular fossils of an early earth-generation by prebiotic methylations and carbamoylations. Angew Chem Int Ed Engl 57(20):5943–5946. https://doi.org/10.1002/anie.201801919. (Epub 17 April 2018)
Shimoyama A, Ogasawara R (2002) Dipeptides and diketopiperazines in the Yamato-791198 and Murchison carbonaceous chondrites. Orig Life Evol Biosph 32(2):165–179. https://doi.org/10.1023/a:1016015319112
Sousa FL, Thiergart T, Landan G, Nelson-Sathi S, Pereira IAC et al (2013) Early bioenergetic evolution. Philos Trans R 368(1622):20130088. https://doi.org/10.1098/rstb.2013.0088. (Print 19 July 2013)
Takeuchi Y (2020) Y Furukawa, T Kobayashi, T Sekine, N Terada & T Kakegawa Impact-induced amino acid formation on Hadean Earth and Noachian Mars. Sci Rep 10:9220
Thornburg ZR, Bianchi DM, Brier TA, Smith HO, Glass JI et al (2022) Fundamental behaviors emerge from simulations of a living minimal cell. Cell 185(2):345–360. https://doi.org/10.1016/j.cell.2021.12.025
Torres de Farias S, José MV (2020) Transfer RNA: The molecular demiurge in the origin of biological systems. Prog Biophys Mol Biol 153:28–34. https://doi.org/10.1016/j.pbiomolbio.2020.02.006
Trifonov EN (2004) The triplet code from first principles. J Biomol Struct Dyn 22:1–11
Trivedi M, Saxena D, Ng WK et al (2022) Self-organized lasers from reconfigurable colloidal assemblies. Nat Phys 18:939–944. https://doi.org/10.1038/s41567-022-01656-2
Wanjura CC, Gago P, Matsushima T, Blumenfeld R (2019) Structural evolution of granular systems: theory. arXiv:1904.06549 (Submitted on 13 April 2019)
Wein T, Sorek R (2022) Bacterial origins of human cell-autonomous innate immune mechanisms. Nat Rev Immunol 22:629–638. https://doi.org/10.1038/s41577-022-00705-4
Wieczorek R, Dörr M, Chotera A, Luisi PL, Monnard P-A (2013) Formation of RNA phosphodiester bond by histidine-containing dipeptides. ChemBioChem 14(2):217–223. https://doi.org/10.1002/cbic.201200643
Wochner A, Attwater J, Coulson A, Holliger P (2011) Ribozyme-catalyzed transcription of an active ribozyme. Science 332(6026):209–212. https://doi.org/10.1126/science.1200752
Wolf YI, Koonin EV (2007) On the origin of the translation system and the genetic code in the RNA world by means of natural selection, exaptation, and subfunctionalization. Biol Direct 2:14. https://doi.org/10.1186/1745-6150-2-14
Wołos A, Roszak R, Żądło-Dobrowolska A, Beker W, Mikulak-Klucznik B (2020) Synthetic connectivity, emergence, and self-regeneration in the network of prebiotic chemistry. Science 369(6511):eaaw1955. https://doi.org/10.1126/science.aaw1955
Xu J, Chmela V, Green N et al (2020) Selective prebiotic formation of RNA pyrimidine and DNA purine nucleosides. Nature 582:60–66. https://doi.org/10.1038/s41586-020-2330-9
Yarus M (1998) Amino acids as RNA ligands: a direct-RNA-template theory for the code’s origin. J Mol Evol 47(1):109–117
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
The authors equally contributed to: study concept and design, acquisition of data, analysis and interpretation of data, drafting of the manuscript, critical revision of the manuscript for important intellectual content, statistical analysis, obtained funding, administrative, technical, and material support, study supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any known or potential conflict of interest including any financial, personal or other relationships with other people or organizations within three years of beginning the submitted work that could inappropriately influence, or be perceived to influence, their work.
Ethical Approval
This research does not contain any studies with human participants or animals performed by the authors.
Consent for Publication
The authors transfer all copyright ownership, in the event the work is published. The undersigned authors warrant that the article is original, does not infringe on any copyright or other proprietary right of any third part, is not under consideration by another journal, and has not been previously published.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tozzi, A., Mazzeo, M. The First Nucleic Acid Strands May Have Grown on Peptides via Primeval Reverse Translation. Acta Biotheor 71, 23 (2023). https://doi.org/10.1007/s10441-023-09474-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10441-023-09474-6